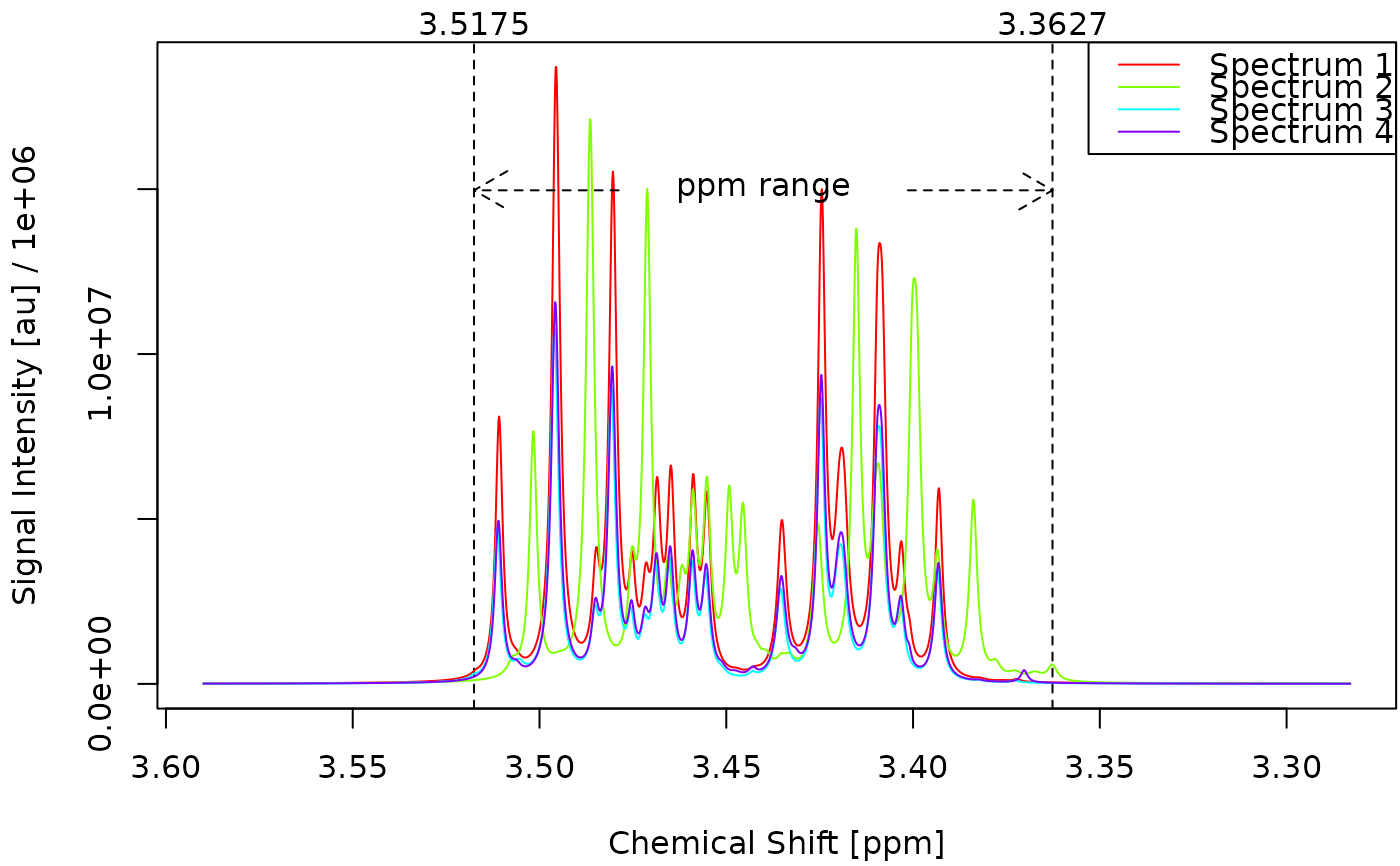
Plot a set of deconvoluted spectra.
Arguments
- obj
An object of type
decons0,decons1ordecons2. For details see Metabodecon Classes.- ...
Additional arguments passed to the conversion function.
- foc_rgn
Numeric vector of length 2 specifying the focus region in ppm (e.g.
c(3.55, 3.35)). If NULL (default), the full spectrum is shown.- what
Which signal to plot:
"supal"(aligned superposition, default with fallback to"sup"then"si"),"sup"(superposition) or"si"(raw).- sfy
Scaling factor for the y-axis.
- cols
Character vector of colors, one per spectrum. Defaults to
rainbow(n).- names
Character vector of legend labels. Defaults to spectrum names.
- xlab
Label for the x-axis.
- ylab
Label for the y-axis.
- mar
A numeric vector of length 4, which specifies the margins of the plot.
- lgd
Logical or list. If TRUE, a legend is drawn at "topright" with
cex = 0.8. If a list, its elements are passed tolegend()to override position, size, etc. Setshow = FALSEinside the list (or passlgd = FALSE) to hide.
See also
plot_spectrum() for a much more sophisticated plotting routine
suitable for plotting a single spectrum.
Examples
obj <- deconvolute(sim[1:4], sfr = c(3.55, 3.35))
#> 2026-04-10 07:15:54.62 Starting deconvolution of 4 spectra using 1 worker
#> 2026-04-10 07:15:54.62 Starting deconvolution of sim_01 using R (legacy) backend
#> 2026-04-10 07:15:54.62 Removing water signal
#> 2026-04-10 07:15:54.62 Removing negative signals
#> 2026-04-10 07:15:54.62 Smoothing signals
#> 2026-04-10 07:15:54.63 Starting peak selection
#> 2026-04-10 07:15:54.63 Detected 314 peaks
#> 2026-04-10 07:15:54.63 Removing peaks with low scores
#> 2026-04-10 07:15:54.63 Removed 287 peaks
#> 2026-04-10 07:15:54.63 Initializing Lorentz curves
#> 2026-04-10 07:15:54.63 MSE at peak tiplet positions: 4.0838805770844048836921
#> 2026-04-10 07:15:54.63 Refining Lorentz Curves
#> 2026-04-10 07:15:54.63 MSE at peak tiplet positions: 0.1609359876216345797140
#> 2026-04-10 07:15:54.64 MSE at peak tiplet positions: 0.0228015051613790313556
#> 2026-04-10 07:15:54.64 MSE at peak tiplet positions: 0.0071638016610617799920
#> 2026-04-10 07:15:54.64 Formatting return object as decon2
#> 2026-04-10 07:15:54.64 Finished deconvolution of sim_01
#> 2026-04-10 07:15:54.64 Starting deconvolution of sim_02 using R (legacy) backend
#> 2026-04-10 07:15:54.64 Removing water signal
#> 2026-04-10 07:15:54.64 Removing negative signals
#> 2026-04-10 07:15:54.64 Smoothing signals
#> 2026-04-10 07:15:54.65 Starting peak selection
#> 2026-04-10 07:15:54.65 Detected 316 peaks
#> 2026-04-10 07:15:54.65 Removing peaks with low scores
#> 2026-04-10 07:15:54.65 Removed 286 peaks
#> 2026-04-10 07:15:54.65 Initializing Lorentz curves
#> 2026-04-10 07:15:54.65 MSE at peak tiplet positions: 3.8338943428876719465848
#> 2026-04-10 07:15:54.65 Refining Lorentz Curves
#> 2026-04-10 07:15:54.65 MSE at peak tiplet positions: 0.1289481941626757499630
#> 2026-04-10 07:15:54.68 MSE at peak tiplet positions: 0.0135651899090413786964
#> 2026-04-10 07:15:54.68 MSE at peak tiplet positions: 0.0025556755331531087749
#> 2026-04-10 07:15:54.68 Formatting return object as decon2
#> 2026-04-10 07:15:54.68 Finished deconvolution of sim_02
#> 2026-04-10 07:15:54.68 Starting deconvolution of sim_03 using R (legacy) backend
#> 2026-04-10 07:15:54.68 Removing water signal
#> 2026-04-10 07:15:54.68 Removing negative signals
#> 2026-04-10 07:15:54.68 Smoothing signals
#> 2026-04-10 07:15:54.69 Starting peak selection
#> 2026-04-10 07:15:54.69 Detected 333 peaks
#> 2026-04-10 07:15:54.69 Removing peaks with low scores
#> 2026-04-10 07:15:54.69 Removed 308 peaks
#> 2026-04-10 07:15:54.69 Initializing Lorentz curves
#> 2026-04-10 07:15:54.69 MSE at peak tiplet positions: 1.4917065120183623516681
#> 2026-04-10 07:15:54.69 Refining Lorentz Curves
#> 2026-04-10 07:15:54.69 MSE at peak tiplet positions: 0.0569971157280155932279
#> 2026-04-10 07:15:54.70 MSE at peak tiplet positions: 0.0065629979536274852397
#> 2026-04-10 07:15:54.70 MSE at peak tiplet positions: 0.0013913916281140725414
#> 2026-04-10 07:15:54.70 Formatting return object as decon2
#> 2026-04-10 07:15:54.70 Finished deconvolution of sim_03
#> 2026-04-10 07:15:54.70 Starting deconvolution of sim_04 using R (legacy) backend
#> 2026-04-10 07:15:54.70 Removing water signal
#> 2026-04-10 07:15:54.70 Removing negative signals
#> 2026-04-10 07:15:54.70 Smoothing signals
#> 2026-04-10 07:15:54.71 Starting peak selection
#> 2026-04-10 07:15:54.71 Detected 325 peaks
#> 2026-04-10 07:15:54.71 Removing peaks with low scores
#> 2026-04-10 07:15:54.71 Removed 299 peaks
#> 2026-04-10 07:15:54.71 Initializing Lorentz curves
#> 2026-04-10 07:15:54.71 MSE at peak tiplet positions: 2.2382155282476525748336
#> 2026-04-10 07:15:54.71 Refining Lorentz Curves
#> 2026-04-10 07:15:54.71 MSE at peak tiplet positions: 0.0843491698981613247099
#> 2026-04-10 07:15:54.71 MSE at peak tiplet positions: 0.0101688144550079444944
#> 2026-04-10 07:15:54.71 MSE at peak tiplet positions: 0.0031861616084395915326
#> 2026-04-10 07:15:54.72 Formatting return object as decon2
#> 2026-04-10 07:15:54.72 Finished deconvolution of sim_04
#> 2026-04-10 07:15:54.72 Finished deconvolution of 4 spectra in 0.095 secs
plot_spectra(obj)